- Blog
- The playroom ps4 prix
- Another thand coming bass songster
- Onedari sharemate 3 p scene
- Frog dissection worksheet answer key pdf
- Flower primordia
- Detroit diesel fire commander ii
- Gentle reader meaning
- 4e shadowrun character sheet
- The hand of fate quest wow
- Capacitor charge time calc
- Ntouch kawasaki adapter
- Luca balsa
- Waterville toast express
- Blog
- The playroom ps4 prix
- Another thand coming bass songster
- Onedari sharemate 3 p scene
- Frog dissection worksheet answer key pdf
- Flower primordia
- Detroit diesel fire commander ii
- Gentle reader meaning
- 4e shadowrun character sheet
- The hand of fate quest wow
- Capacitor charge time calc
- Ntouch kawasaki adapter
- Luca balsa
- Waterville toast express
(H) XTH9 expression is down-regulated in 35S:ALCR/AlcA:ANT-IR and 35S:ALCR/AlcA:AIL6-amiR ant inflorescences after ethanol treatment. (G) AN3 expression is slightly down-regulated in 35S:ALCR/AlcA:ANT-IR inflorescences and more severely down-regulated in 35S:ALCR/AlcA:AIL6-amiR ant inflorescences. (F) ROT3 expression is down-regulated in 35S:ALCR/AlcA:ANT-IR and 35S:ALCR/AlcA:AIL6-amiR ant inflorescences after ethanol treatment. (E) BB expression is up-regulated in 35S:ALCR/AlcA:ANT-IR and 35S:ALCR/AlcA:AIL6-amiR ant inflorescences after ethanol treatment. (D) Expression of ANT is reduced in 35S:ALCR/AlcA:ANT-IR inflorescences after a 24 h EtOH treatment. (C) Stamen anther of a mock-treated (left) and ethanol-treated (right) 35S:ALCR/AlcA:ANT-IR flower. Between 10 and 20 petals from different flowers were measured for each genotype and treatment. (B) Petal area, length, and width measurements for mock- and EtOH-treated L er and 35S:ALCR/AlcA:ANT-IR flowers. (A) Mock- and ethanol (EtOH)-treated 35S:ALCR/AlcA:ANT-IR flowers. Published by Oxford University Press on behalf of the Society for Experimental Biology.Įxpression of growth-regulatory genes is altered after down-regulation of ANT expression in 35S:ALCR/AlcA:ANT-IR inflorescences and after down-regulation of AIL6 expression in 35S:ALCR/AlcA:AIL6-amiR ant inflorescences. These include class B and C floral homeotic genes, growth regulatory genes, and genes involved in vascular development.Īrabidopsis thaliana AINTEGUMENTA (ANT) AINTEGUMENTA-LIKE6 (AIL6) ChIP-Seq floral homeotic genes floral organ identity flower development organ growth organ size vascular development. Comparison of genes associated with both ANT and AIL6 ChIP-Seq peaks and those differentially expressed after perturbation of ANT and/or AIL6 activity identified likely direct targets of ANT and AIL6 regulation. ANT- and AIL6-binding sites are associated with genes involved in many biological processes related to meristem and flower organ development. AIL6 binds to a subset of ANT sites, suggesting that AIL6 regulates some but not all of the same target genes as ANT. To gain insight into the molecular mechanisms by which ANT and AIL6 contribute to floral organogenesis, we identified the genome-wide binding sites of both ANT and AIL6 in stage 3 flower primordia, the developmental stage at which sepal primordia become visible and class B and C floral homeotic genes are first expressed.

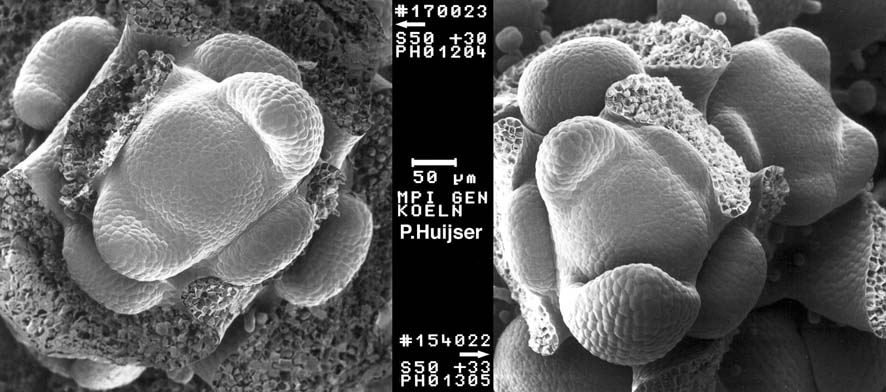
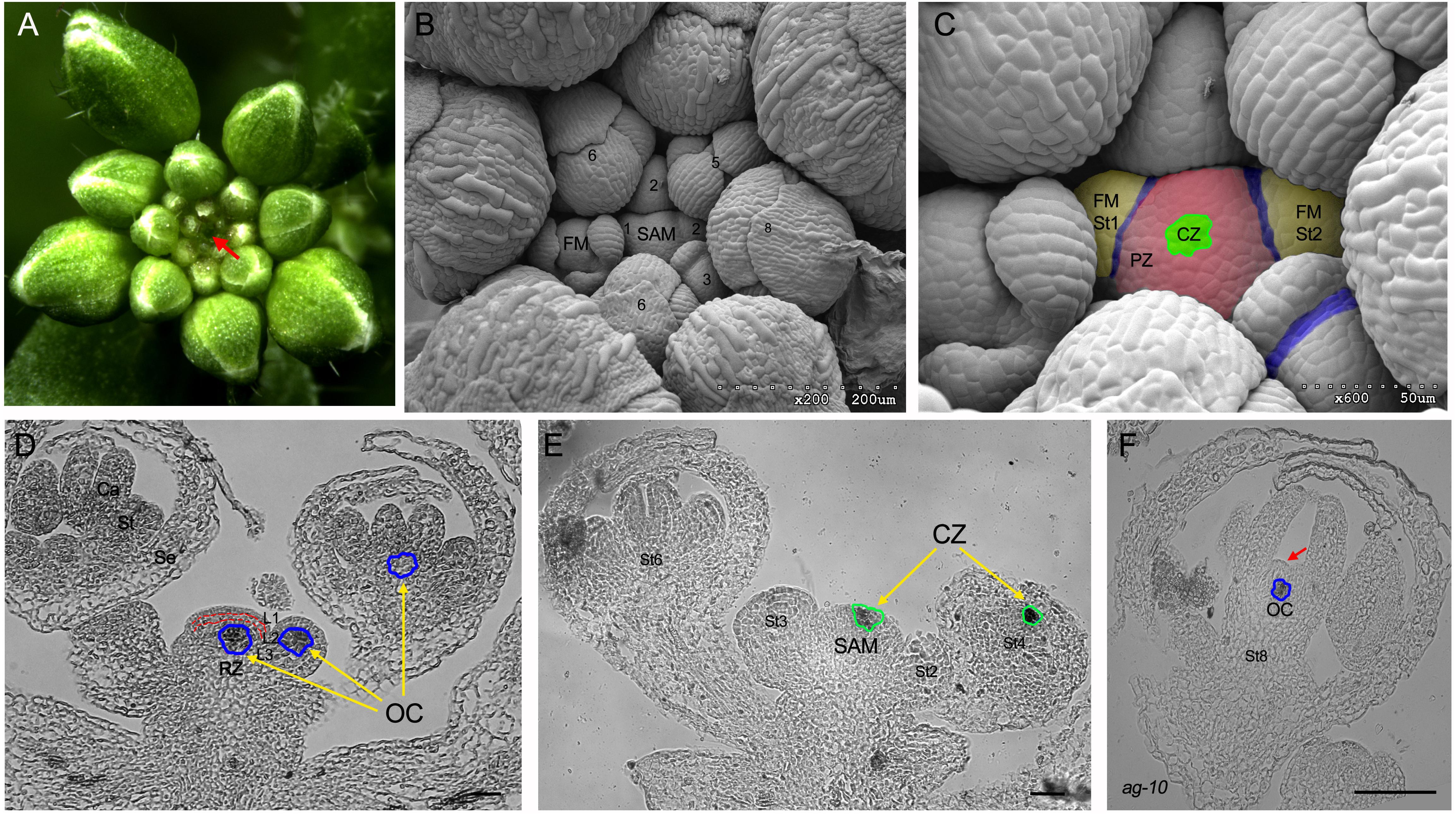
The Arabidopsis transcription factors AINTEGUMENTA (ANT) and AINTEGUMENTA-LIKE6 (AIL6) are required for correct positioning of floral organ initiation, contribute to the specification of floral organ identity, and regulate the growth and morphogenesis of developing floral organs. In the control, only 11 of haploids produced selfed kernels owing to spontaneous chromosome doubling. Floral organ primordia in each whorl undergo distinct developmental programs to become one of four organ types (sepals, petals, stamens, and carpels). Treatment at the sixleaf stage (flower primordia formation stage) significantly increased the occurrence of fertile sectors on both tassels and ears so that approximately half (44) of the treated haploids produced kernels after selfpollination. Arabidopsis flower primordia give rise to organ primordia in stereotypical positions within four concentric whorls.
